Reading the Book of Life: How Advanced DNA Sequencing Is Transforming Medicine

We all know that DNA (four chemical letters: A, T, C, G) encodes the genetic information of our lives, including appearance, growth, and even genetic diseases. It performs like an instruction book that tells our cells how to function. However, do you know how we decrypt this personal book and apply it for diagnosis and the development of therapies? Today, thanks to powerful sequencing technologies, scientists can read entire genomes faster than ever before, transforming biomedical research and modern medicine.
The first generation of DNA sequencing methods started from Sanger sequencing, which is like reading a book one page at a time and stopping at specific letters to see where they occur. This makes it precise, but too slow for reading large genomes. Then, next-generation sequencing (NGS) occurs, which could sequence millions of reads simultaneously. It performs like tearing a book into thousands of small pieces, reading them all at once, and then using a computer to put the story back together. However, the process is complex, and the equipment used is large and expensive, making it unsuitable for remote areas.
More recently, nanopore sequencing has pushed the boundaries. This technology reads DNA by passing a single strand through a tiny pore and measuring changes in electrical current1. Remarkably, nanopore sequencers can be small enough to fit in the palm of your hand. This portability means DNA sequencing is no longer confined to large, well-funded laboratories — it can be used in hospitals, field sites, and even low-resource settings.

Figure 2. The evolution of DNA sequencing technology. Figure generated from GPT-5.
You might think DNA is boring—a long, unchanging string made up of just A, T, C, and G. But DNA is much cooler than that: it’s constantly dynamic, changing, and adapting every second. For instance, chemical modifications can occur on certain nucleotides, such as methylation or phosphorylation. Think of these like little notes you write in the margins of a book to help you remember important points when you review it. On top of that, mistakes can happen during “printing,” which might confuse your reading—this is what we call DNA damage.
In this context, nanopore sequencing could also help identify these sites and reveal their distribution across the genome. For example, an abasic site is a position in DNA where a base is missing2. It could lead to incorrect replication or transcription in that region, causing diseases and cancer. As a first-year PhD student, my first rotation project involves training a model to distinguish it from the other 4 normal nucleotides in Professor Martin Taylor’s lab. What’s exciting is that this project combines hands-on lab work with data analysis and coding. The library was prepared by synthesizing oligos that have an abasic site in the middle. After sequencing, we can compare the nanopore signal differences to build a model that accurately detects these damaged sites (Figure 3).
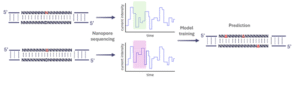
Figure 3. The workflow of training nanopores to detect abasic sites.
By establishing this model, nanopore sequencing can be used to explore how abasic sites contribute to disease and how cells repair them. Beyond basic research, this approach also has translational potential for diagnostics and identifying therapeutic targets.
References:
1 Wang, Y., Zhao, Y., Bollas, A., Wang, Y. & Au, K. F. Nanopore sequencing technology, bioinformatics and applications. Nature Biotechnology 39, 1348-1365 (2021). https://doi.org/10.1038/s41587-021-01108-x
2 Thompson, P. S. & Cortez, D. New insights into abasic site repair and tolerance. DNA Repair (Amst) 90, 102866 (2020). https://doi.org/10.1016/j.dnarep.2020.102866



